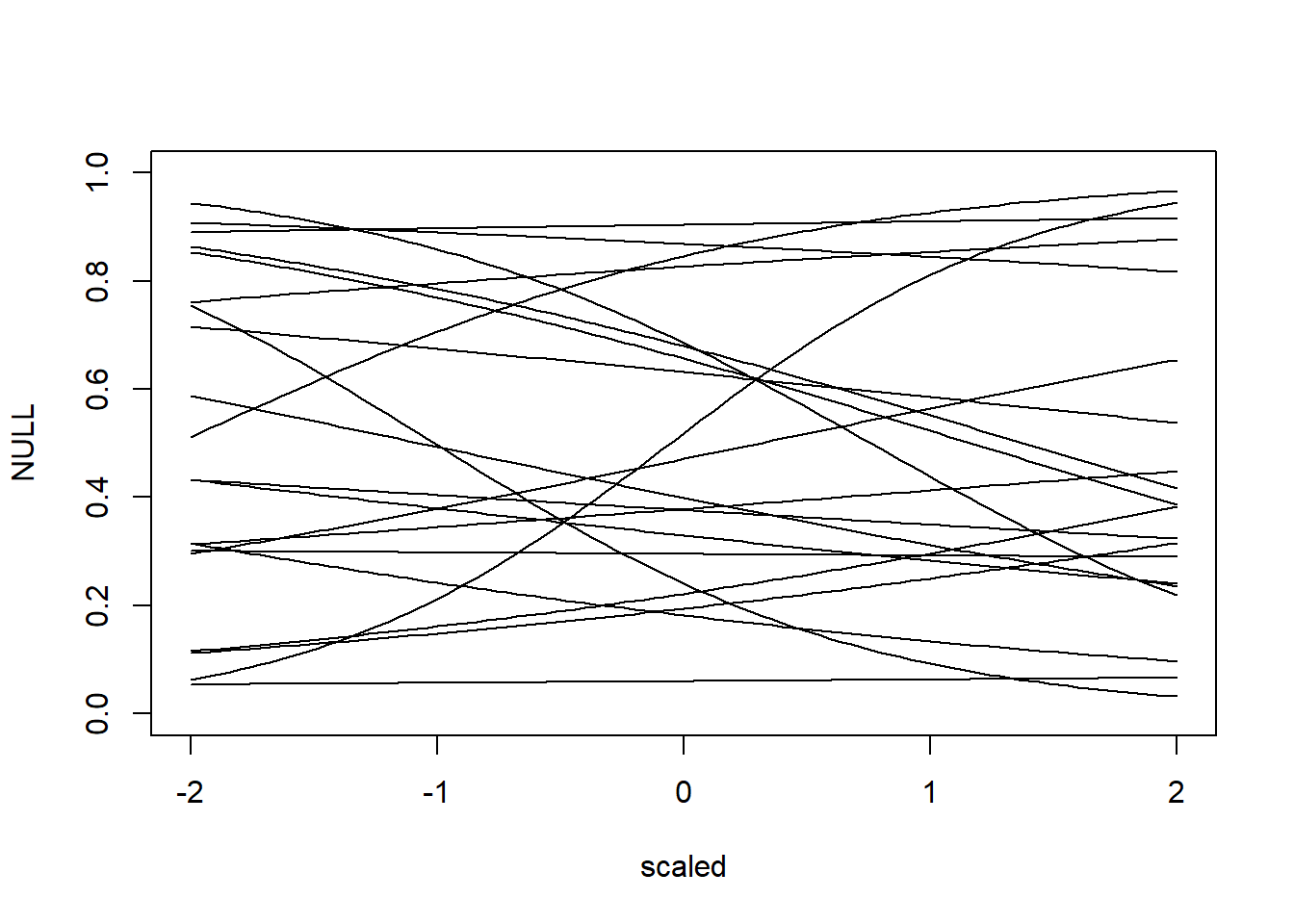
N <- 20 # number of simulations
alpha <- qlogis(rbeta(N, 2, 2))
beta <- rnorm(N, 0, 0.5)
plot(NULL, xlim = c(-2, 2), ylim = c(0, 1), xlab = "scaled")
for (i in 1:N) {
curve(plogis(alpha[i] + beta[i] * x), from = -2, to = 2, add = TRUE)
}S1 Data preparation
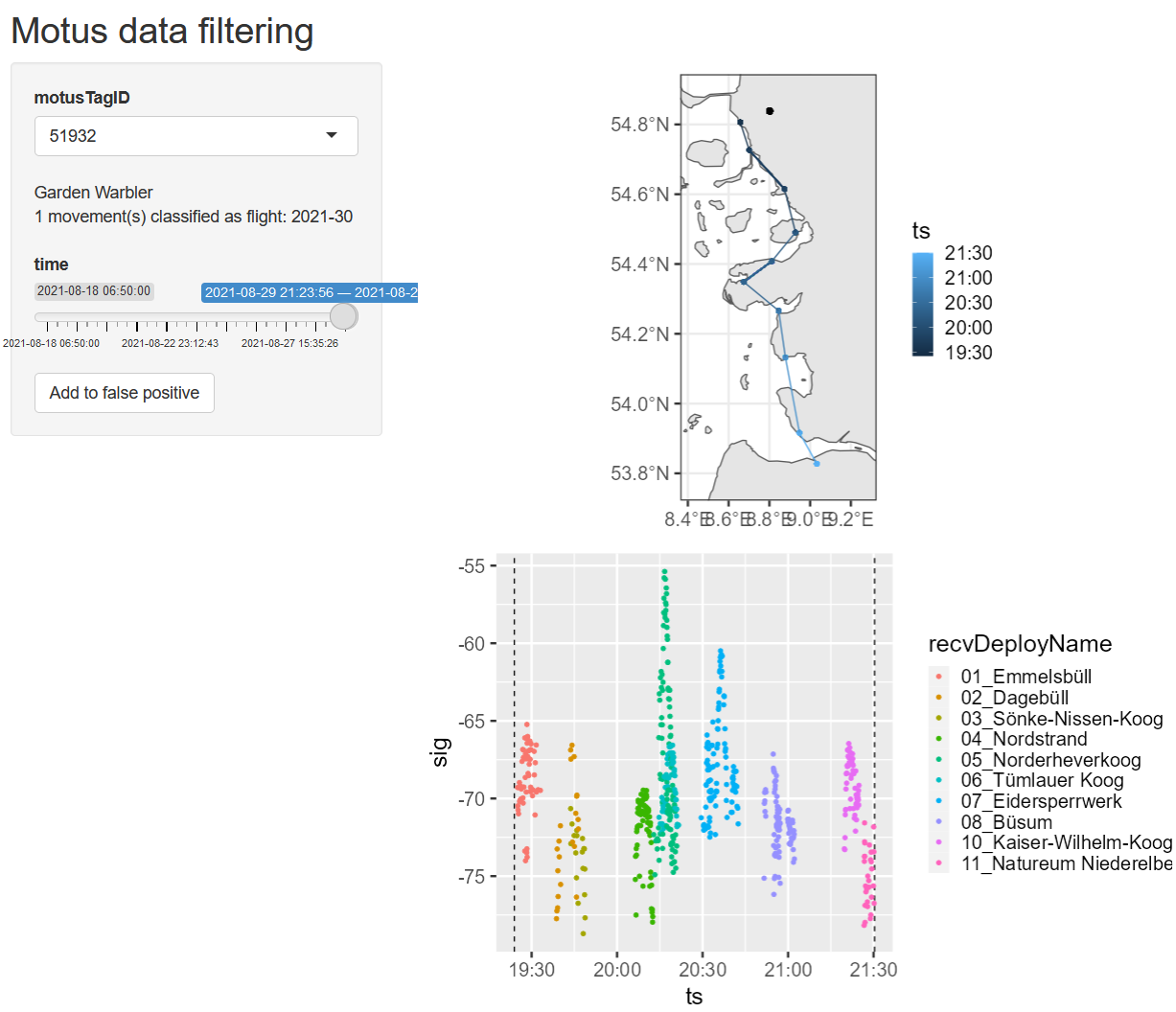
Data filtering
runLen> 3- exclude detections in the middle of nowhere
- remove manually classified false positives
Flight parameter determination
NoteFlight definition
Starting east of 8°E and north of 54°N with a distance of > 50 km between first and last receiver.
flightIDflight identifierflightStartstart time (max signal strength at first receiver within last half hour)flightEndend time (last detection)flightDatesunset date of departureminStopoverminimum stopover duration in days
Find departures without detected flights: movements of > 50 km within one day.
minLengthminimum movement length
Validate data
Run Shiny app data/app.R to:
- manually remove false positives
- validate flight parameters
Flight classification
S2 Simulation
Prior predictive simulations
Slopes
In [1]:
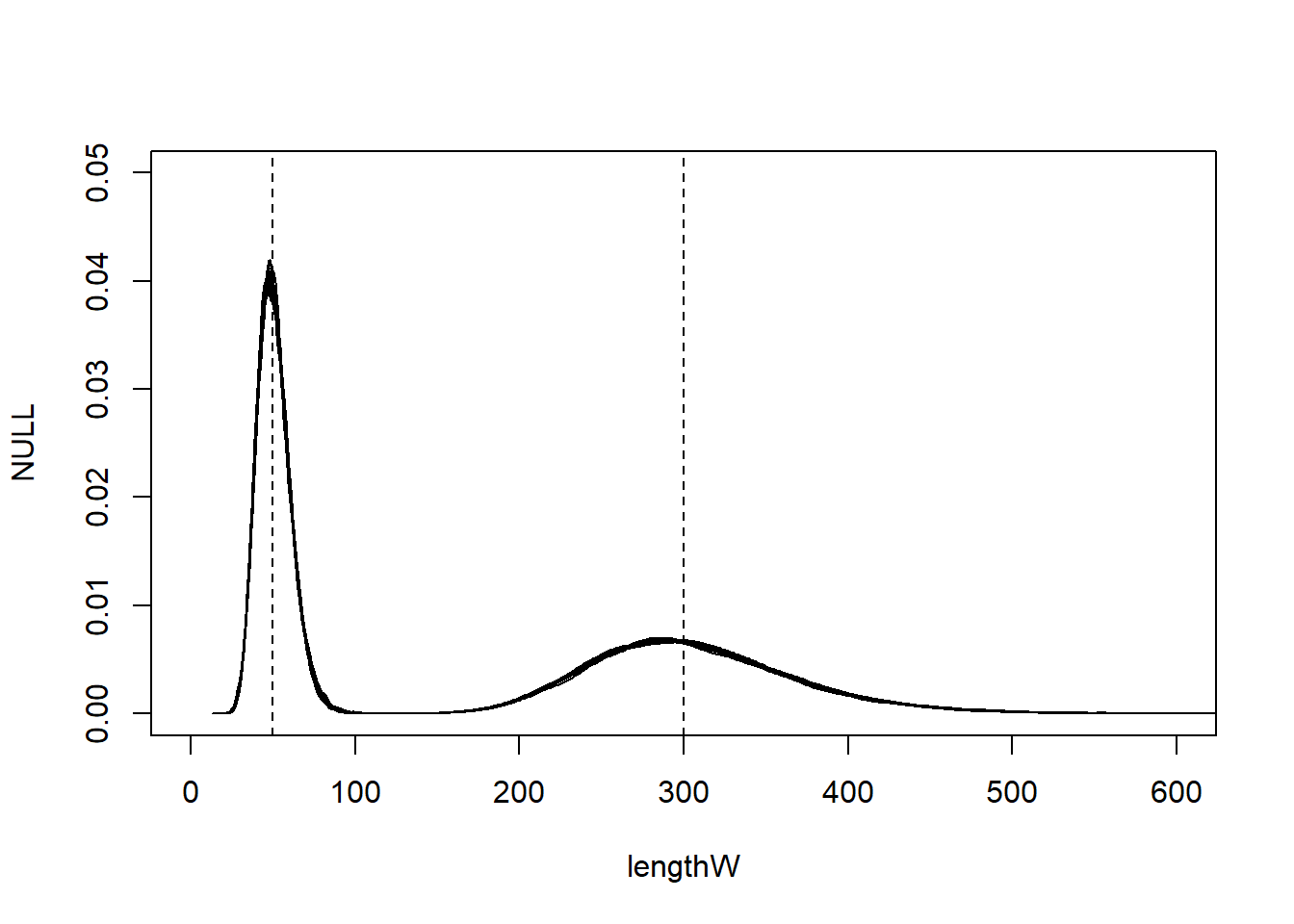
Measurement error
Generative model
Note
The code to simulate and analyse data from the generative model can be found in 4_mixture_simulation.R.